Mlipaudit Benchmarks ML Interatomic Potentials for Precise, Efficient Simulations
The growing need for accurate yet efficient atomistic simulations has accelerated the development of machine-learned interatomic potentials (MLIPs), which can model complex molecular systems at far lower computational cost than traditional electronic structure methods. Despite this potential, the field has lacked standardised tools for evaluating and comparing MLIP performance across diverse chemical environments.
To overcome this gap, Leon Wehrhan, Lucien Walewski, Marie Bluntzer, and colleagues at InstaDeep have developed MLIPAudit, an open, modular, and comprehensive benchmarking suite. MLIPAudit systematically assesses MLIP accuracy across a wide range of tasks—including organic molecules, liquids, proteins, and peptides—and provides an actively updated leaderboard for transparent model comparison. By offering curated benchmark datasets and pre-computed results for numerous publicly available MLIPs, the framework establishes a unified, reproducible standard for validation. This effort is expected to streamline MLIP discovery, foster comparability, and accelerate progress toward reliable, broadly applicable molecular simulation tools.

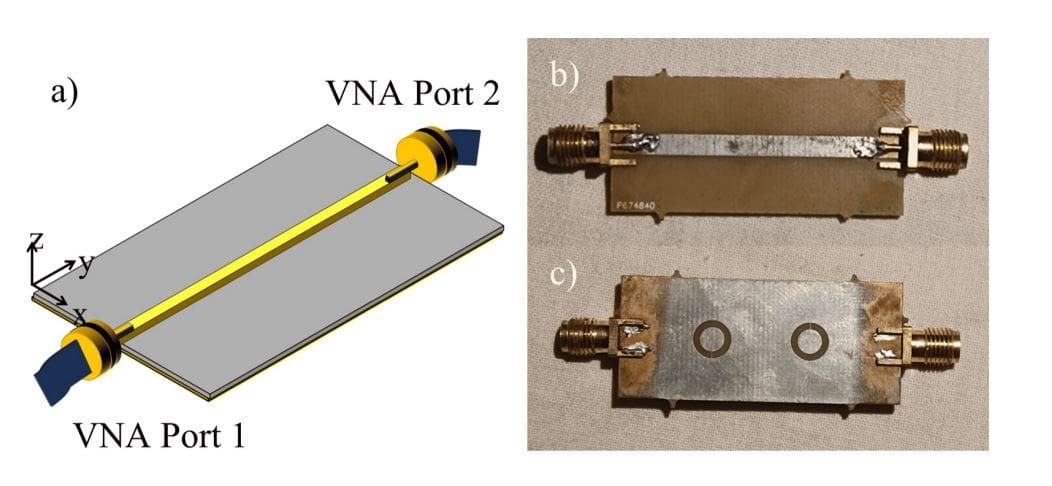
Figure 1. MLIPAudit Evaluates Machine-Learned Interatomic Potentials
Validation Datasets for Machine-Learning Interatomic Potentials
The rise of machine learning potentials (MLPs) as alternatives to conventional force fields has spurred the development of diverse datasets and software tools for their training, validation, and deployment. Key resources include SPICE for drug-like molecules and peptides, Transition1x for reactive systems, and Wiggle150 for strained conformers that challenge model robustness. Universal pre-trained models such as CHGNet, along with extensive quantum-chemistry–derived datasets of reactants, products, and transition states, further support MLP development. A large dataset of 134,000 molecules also aids quantum structure analysis. Figure 1 shows MLIPAudit Evaluates Machine-Learned Interatomic Potentials.
Benchmarking frameworks like MLIPAudit enable standardized evaluation, while tools such as ORCA provide methods like the Nudged Elastic Band (NEB) for transition-state identification. Popular MLP approaches—including ANI-1 and TorsionNet, which predicts torsional energy profiles—are increasingly adopted. Meanwhile, traditional force fields continue to evolve through initiatives such as the Open Force Field (OFF) project.
Versions 0 and 2 of the Open Force Field project reflect an ongoing collaborative effort to develop a broadly applicable, next-generation force field. Established frameworks such as the Amber Force Field and the General Amber Force Field (GAFF) continue to support molecular dynamics simulations, while a range of widely used water models—SPC/E, TIP3P, TIP4P, TIP5P, and the Jorgensen model—remain essential for accurately representing aqueous environments. These classical models are complemented by quantum chemistry and electronic structure approaches, including Density Functional Theory (DFT) and the Nudged Elastic Band (NEB) method, which supply high-quality reference data for training and validating machine learning potentials.
Molecular dynamics tools and analytical techniques, such as calculating radial distribution functions (RDFs) for liquids like water, carbon tetrachloride, methanol, and acetonitrile, play a key role in characterizing molecular structure. Extensive datasets—featuring experimental diffraction data and computed RDFs for water and other liquids—provide critical benchmarks. Additional resources, including Tautobase for tautomer enumeration and SPICE for drug-like molecules and peptides, support specialized applications.
Overall, the field is transitioning toward machine learning potentials, emphasizing rigorous benchmarking, improved water modeling, and achieving DFT-level accuracy with significantly reduced computational cost—an evolution driven by open science and collaborative innovation./p>
MLIPAudit Evaluates Interatomic Potential Accuracy
Scientists have introduced MLIPAudit, a comprehensive benchmarking suite designed to rigorously evaluate the performance of machine-learned interatomic potentials (MLIPs). Addressing a critical gap in the field, MLIPAudit moves beyond traditional energy and force error metrics to assess model stability, transferability, and real-world applicability. The framework provides a standardized approach for benchmarking MLIPs across diverse systems, including small organic molecules, molecular liquids, and biomolecules.
MLIPAudit supplies curated reference datasets and tools for systematic validation, fostering reproducibility and transparency. The suite evaluates models not only on energy and force accuracy but also on their ability to predict properties relevant to downstream applications. By incorporating tests of stability, robustness, and transferability, it delivers a holistic assessment of model performance. The researchers demonstrate MLIPAudit’s utility through applications to both internal and publicly available MLIP models, including UMA-Small, MACE-OFF, and MACE-MP.
The data generated by MLIPAudit enables clear comparisons of model performance across diverse benchmarks, supporting informed selection of the most suitable MLIP for specific simulations. Its modular design allows easy expansion and contributions from the broader scientific community. The suite is open-source, available on GitHub and PyPI under the Apache License 2.0, and features a continuously updated leaderboard on HuggingFace to track benchmark results. This collaborative approach fosters transparency, reproducibility, and rapid advancement in the development and deployment of accurate, efficient MLIPs for complex molecular systems.
Standardized Evaluation of Interatomic Potentials
MLIPAudit marks a major advancement in the evaluation of machine-learned interatomic potentials (MLIPs), providing a comprehensive, open-source benchmarking suite for systematic performance assessment. The curated repository includes benchmark systems spanning small molecules, molecular liquids, and biomolecules, addressing the need for standardized and reproducible evaluation protocols. By emphasizing rigorous validation and direct comparison rather than model-centric testing alone, MLIPAudit enables thorough assessment of accuracy, transferability, and robustness across diverse predictive models.
The suite allows researchers to move beyond traditional metrics, such as energy and force errors, toward evaluations that better reflect real-world simulation requirements. While the current benchmarks cover a limited set of systems and properties, future expansions will incorporate a broader range of materials and application scenarios, further enhancing the tool’s utility. Freely accessible via GitHub, PyPI, and HuggingFace, MLIPAudit promotes transparency, collaboration, and accelerated development in the field of machine-learned interatomic potentials.
References:
- https://quantumzeitgeist.com/machine-mlipaudit-benchmarks-learned-interatomic-potentials-accurate-cost-effective/
Cite this article:
Janani R (2025), Mlipaudit Benchmarks ML Interatomic Potentials for Precise, Efficient Simulations, AnaTechMaz, pp. 254